Kit for the determination of the CAG and CTA/CTG triplet repeats number (SCAs 1, 2, 3, 6, 7 and SCA 8, respectively) by fluorescent fragment analysis
Information about the product
The autosomal-dominant spinocerebellar ataxias (ADCA) are a heterogeneous group of neurodegenerative disorders characterized by slowly progressive cerebellar dysfunction.
Over 17 distinct types of hereditary spinocerebellar ataxia (SCA) disorders have been described. Because of variable expression and phenotypic overlap, the SCA disorders cannot be differentiated reliably on a clinical basis; an accurate diagnosis depends on molecular testing that detects a mutation in a specific causative gene.
The most common types of SCAs are 1, 2, 3, 6 and 7 (caused by CAG trinucleotide repeats) and SCA8 (CTA/CTG expansions)
Intended use
Adellgene® SCAs is a semi-automated in vitro diagnostic kit designed for use in clinical laboratories which quantitatively determines the number of repetitions of CAG of the SCA 1, 2, 3, 6, 7 at the beginning of the exons of the corresponding genes, and CTA/CTG repeats in the 3’UTR region in the SCA 8 gene. It aims to aid clinical diagnosis associated with autosomal dominant spinocerebellar ataxia.
The use of this kit is the determination of both healthy and unhealthy alleles which have a size equal or less than 200 repeats. The range of classification depends on the specific SCA type. Large abnormal expansions described for SCA 2, SCA 7, and SCA 8 with more than 200 repeats are not detected for this kit and it is necessary the use of other methodology to solve this kind of samples.
The technology is based on a multiplex polymerase chain reaction (PCR) of genomic DNA extracted from peripheral blood, followed by fluorescence analysis of the size of the PCR fragments obtained by genetic analyzer and conversion of that size in the number of CAG or CTA/CTG repeats.
Patients who can benefit from this determination are those referred by a specialist. The intended user of the kit is technical personnel trained to carry out the protocol and the interpretation of results described in the instructions for use.
Workflow

Results
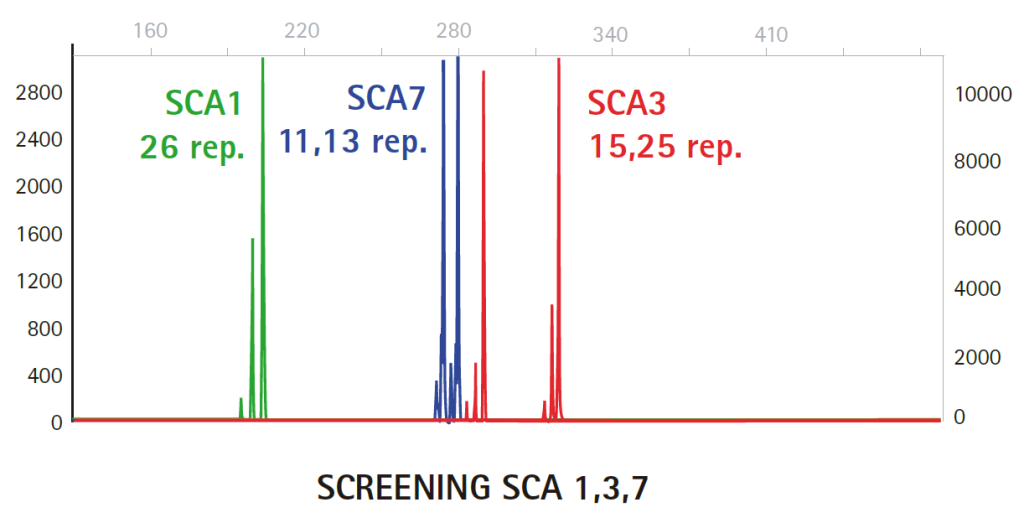
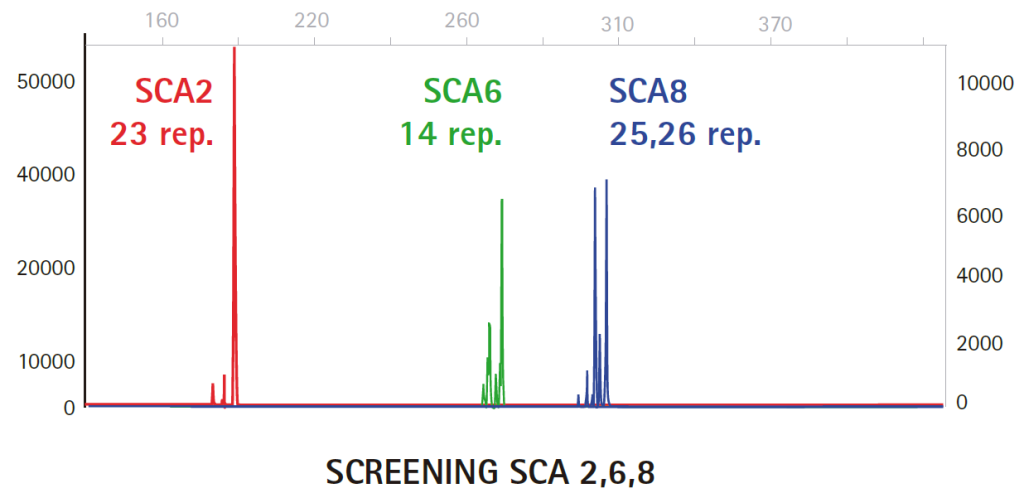
Limitations
- Mutations (point mutations, insertions, deletions) at amplification primer sites are possible and may result in the lack of allele definition. Other technologies could be necessary to resolve the genotyping.
- Homozygous results for SCA 2, 7 and 8 must be confirmed by alternative procedures.
- Data and result interpretation should be revised by qualified personnel.









