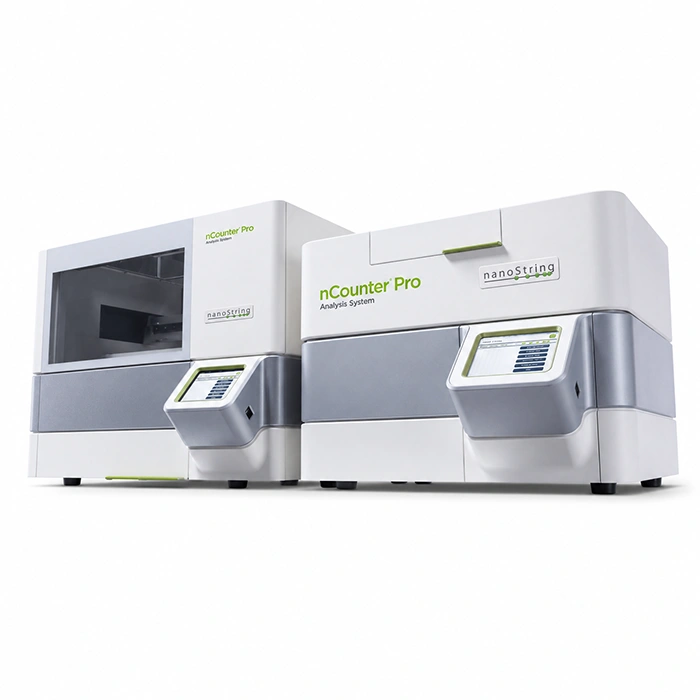
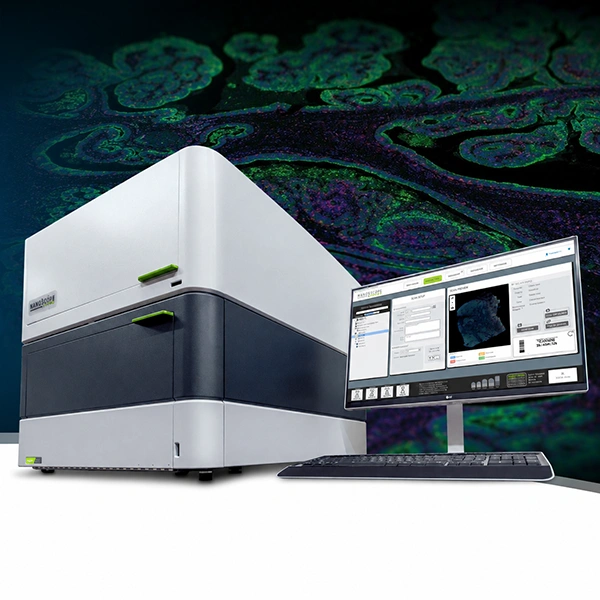
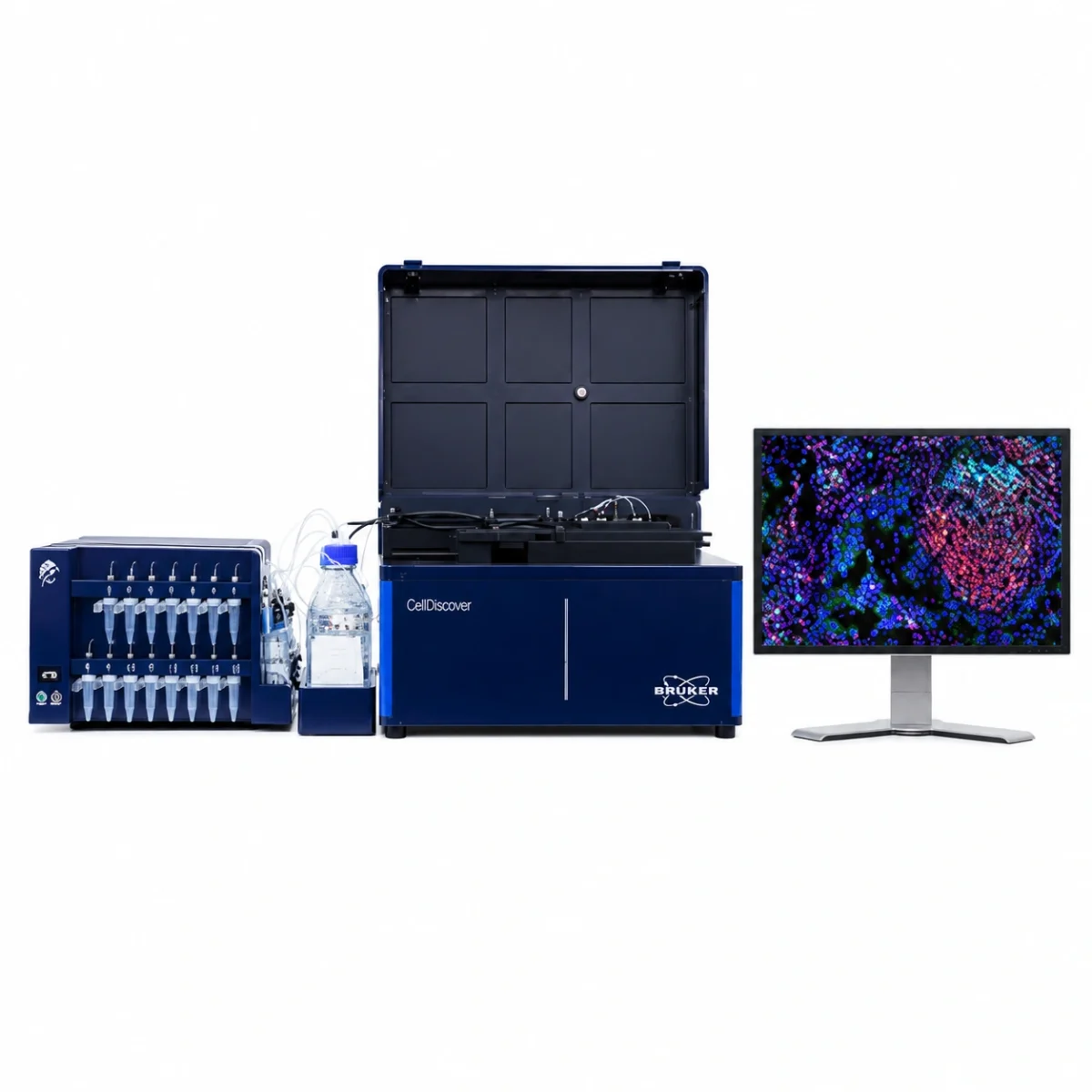
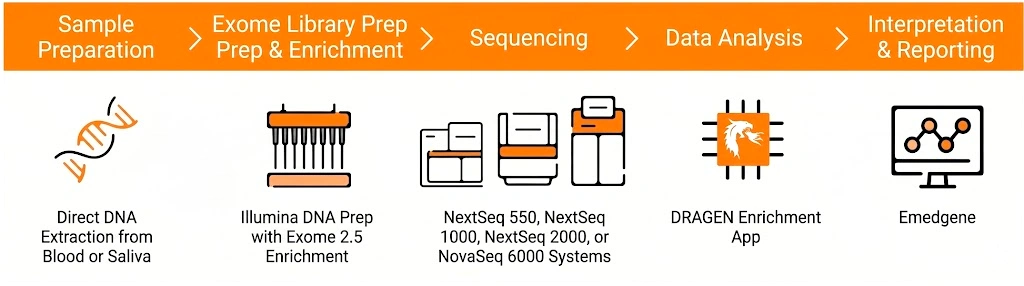
Comprehensive solution for whole exome sequencing with Illumina technology, combining library preparation by tagmentation with capture-based enrichment, sequencing, and bioinformatics analysis, enabling a single workflow.
Detailed Description
Operating Principle
The workflow uses magnetic bead-linked transposomes (eBLT) for rapid and uniform tagmentation. Following index PCR, a hybrid capture enrichment step is performed using the Twist Bioscience for Illumina Exome 2.5 panel. Once the process is complete, the libraries are ready for sequencing. Additionally, secondary and tertiary analyses are seamlessly handled by the DRAGEN™ and Emedgene modules, respectively.
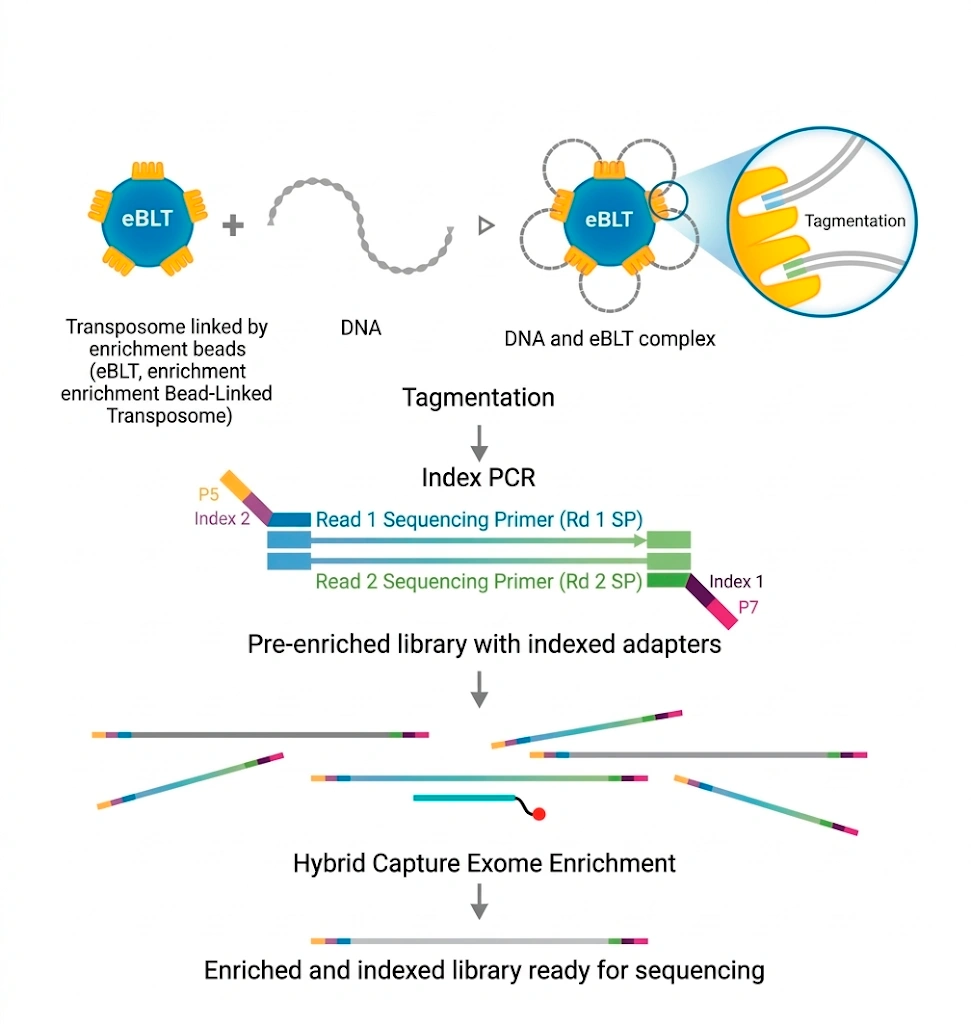
Clinical Applications
Whole exome sequencing (WES) is currently an essential tool in clinical practice for identifying genomic alterations with diagnostic and/or therapeutic relevance. Compared to whole genome sequencing, it offers lower costs and optimal coverage of the coding regions of the human genome, facilitating more precise diagnoses and informed medical decision-making.
Benefits
Intended Audience/User
Molecular diagnostic laboratories working with human genomic DNA samples to diagnose genetic diseases using whole exome techniques.
Key Features
Ordering Information
Kit Components
| 20077595 | Illumina DNA Prep with Exome 2.5 Enrichment, (S) Tagmentation Set B (96 samples, 12‑plex) |
| 20077596 | Illumina DNA Prep with Exome 2.5 Enrichment, (S) Tagmentation Set D (96 samples, 12‑plex) |
| 20091654 | UD Indexes (96 indexes/96 samples): Set A |
| 20091656 | UD Indexes (96 indexes/96 samples): Set B |
| 20091658 | UD Indexes (96 indexes/96 samples): Set C |
| 20091650 | UD Indexes (96 indexes/96 samples): Set D |
Additional Reagents (Extraction from Blood)
| 20018706 | Flex Lysis Reagent Kit (96 reactions). Required for direct input of unextracted peripheral blood. |
Optional Enrichment Panels
| 20093180 | Twist Bioscience for Illumina Mitochondrial Panel (96 samples, 12‑plex). Full coverage of chrM (16,659 bp; 37 genes). |
| Inquire | Illumina Custom Enrichment Panel v2, custom spike-in to expand content or deepen coverage in specific genomic regions. |