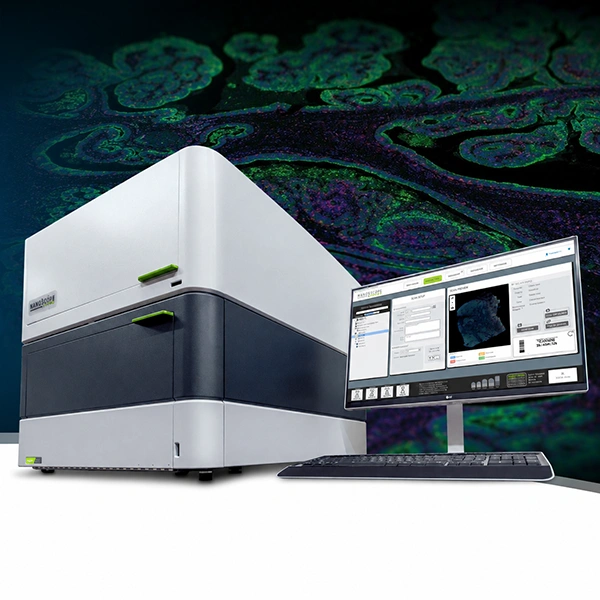
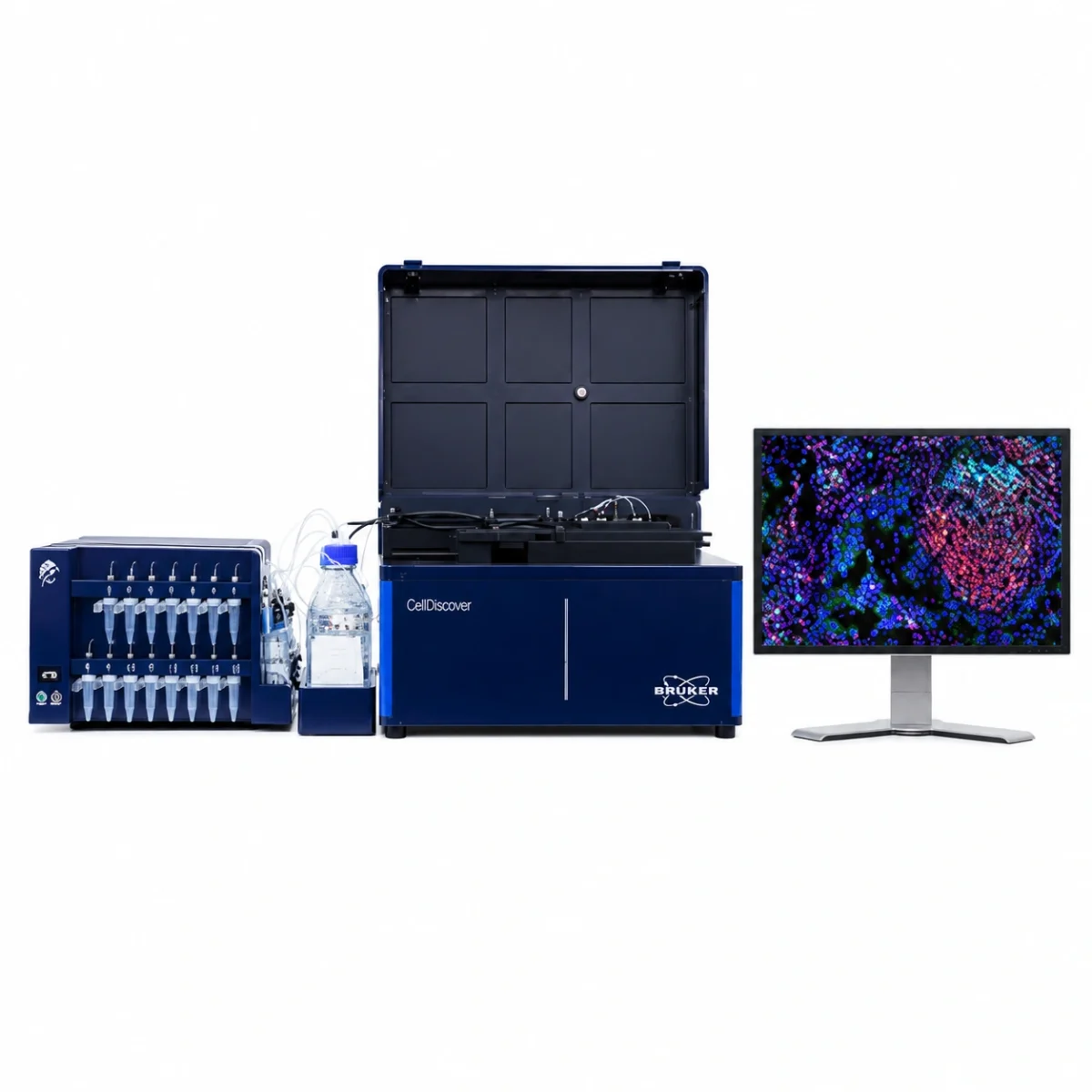
Bruker’s GeoMx™ Digital Spatial Profiler (DSP) is an advanced platform for multi-omic spatial analysis that enables simultaneous quantification of RNA and proteins in tissue sections, preserving tissue architecture. GeoMx DSP combines fluorescent imaging with photo-cleavable oligonucleotide barcodes, offering digitally quantified molecular profiles in defined regions of interest (ROI).
This technology supports FFPE and fresh frozen (FF) samples, and allows the selection of specific tissue areas to obtain expression data with spatial resolution, through an automated and scalable workflow that integrates with standard histological staining techniques. GeoMx DSP offers a high level of multiplexing: from targeted panels to whole transcriptome via NGS, along with over 1,200 validated proteins, providing a comprehensive view of the tissue microenvironment.
With GeoMx DSP, researchers can map the spatial distribution of biomarkers, explore cellular interactions, and characterize complex biological processes in their native context. It is ideal for studies in oncology, immunology, neuroscience, and drug development, facilitating biomarker discovery and a deep understanding of spatial biology.
Key Features
How does it work?
Whole slides are stained with assay probes and up to four fluorescent markers (morphology markers), and subsequently imaged on the instrument. Guided by the morphology markers, the regions of interest (ROI) of the tissue to be profiled are chosen, and by using masks (areas of illumination, AOI), two or more tissue compartments or cell types within each ROI can optionally be profiled. RNA and protein expression levels are quantified by direct digital counting of barcodes using the nCounter® system or by sequencing on an Illumina platform. Data is analyzed using the open-source DSP Data Analysis (DSPDA) suite.
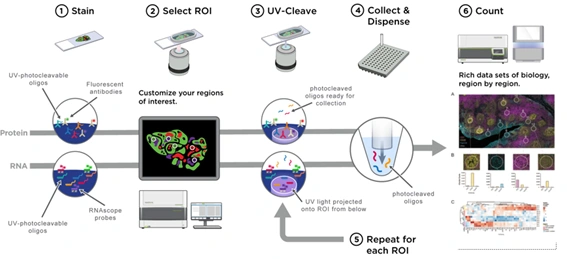
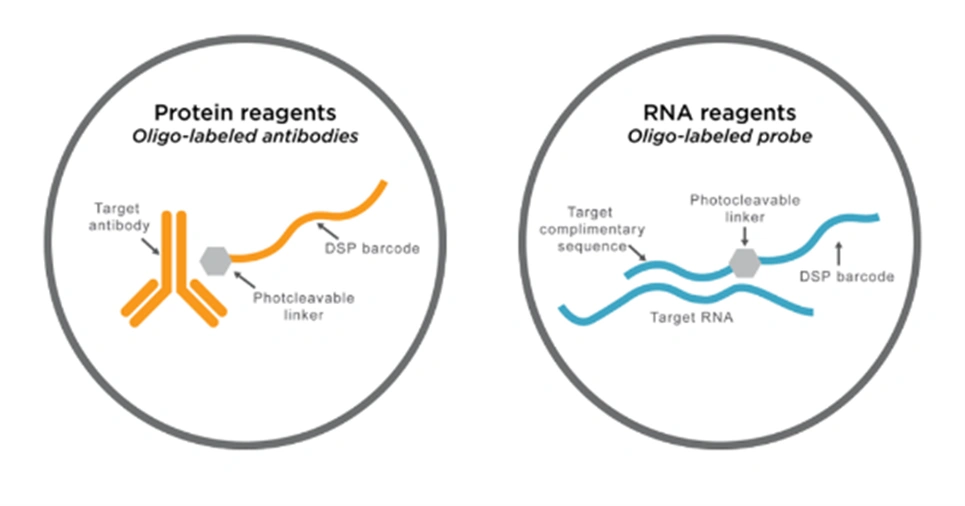
DSP probes are labeled with unique oligonucleotide barcodes that are released upon UV light exposure, without damaging the tissue.
Tissue samples are stained with imaging probes (morphology markers) and fluorophore-labeled assay probes using a standard histological workflow.
Regions of interest (ROIs) and areas of illumination (AOIs) are selected on the tissue for interrogation. The tissue is exposed to UV light, and the barcodes are released and subsequently quantified.
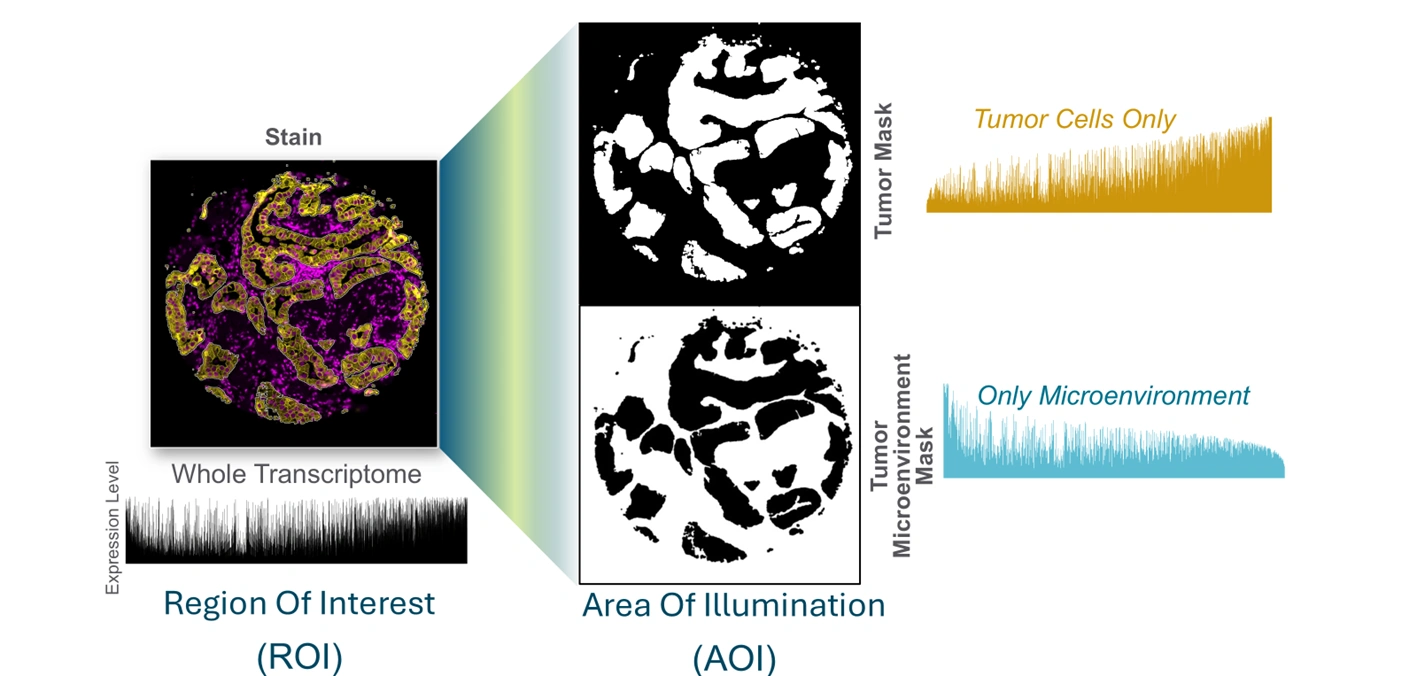
Unbiased spatial expression is obtained from each tissue mask, with the sensitivity to detect low, medium, and high expressors.
Applications
Available panels:
Flexible options to meet your research needs. For more information, click on the corresponding icon:
GeoMx DSP RNA Assays

GeoMx® Human Whole Transcriptome Atlas

GeoMx® Mouse Whole Transcriptome Atlas

GeoMx® Cancer Transcriptome Atlas

GeoMx® TCR Profiling Add-On

GeoMx® Canine Cancer Atlas

GeoMx® Immune Pathways Panel
GeoMx DSP Protein Assays

GeoMx® Discovery Proteome Atlas

GeoMx® ImmunoOncology Proteome Atlas

GeoMx® Immuno-Oncology Protein Panels

GeoMx® Neuroscience Protein Panels









