Kit for the determination of healthy, premutation and mutant alleles in the FMR1 gene by fluorescent fragment analysis
Information about the product
Fragile X Syndrome (FXS, OMIM #300624) is an X-linked disease that is primarily based on the genomic expansion of a triplet of nucleotides (CGG), and aberrant methylation of the promoter region.
FXS has a prevalence of 1 in 4000 males and 1 in 8000 females. Affected individuals show a striking phenotype consisting on large ears and a prominent jaw.
Depending on the number of repetitions of this triplet, four categories can be established:
- Up to 44 repeats: individuals with healthy alleles.
- From 45 to 54 repeats: individuals with intermediate alleles.
- From 55 to 200 repeats: individuals with premutation alleles.
- Over 200 repeats: individuals with mutant alleles.
Intended Use
Adellgene® Fragile X is a semi-automated in vitro diagnostic kit designed for use in clinical laboratories for the amplification and quantitative determination of CGG triplet repeats (cytosine-guanine-guanine) in the 5’ untranslated region of gene for fragile X mental retardation (“Fragile X mental retardation-1”: FMR1). It aims to aid diagnosis of clinical disease associated with Fragile X syndrome (e.g.: mental retardation, primary ovarian failure, tremors / ataxia, etc …).
The technology is based on the polymerase chain reaction (PCR) amplification of genomic DNA extracted from peripheral blood, followed by fluorescence analysis of the size of the PCR fragments obtained by genetic analyzer.
Patients who can benefit from this determination are those referred by a specialist. The intended user of the kit is technical personnel trained to carry out the protocol and the interpretation of results described in the Instructions for Use.
Workflow

Results
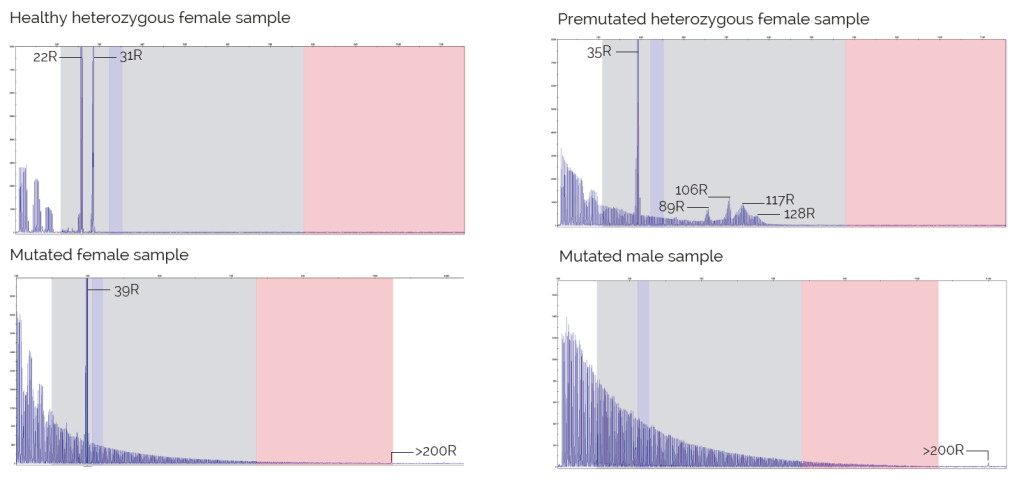
Limitations
- This kit can quantify all alleles up to 200 CGG repeats.
- Mutations (point mutations, insertions, deletions) at amplification primer sites are possible and may result in the lack of allele definition. Other technologies could be necessary to resolve the genotyping.
- Data and result interpretation should be revised by qualified personnel.