Exome designed to provide comprehensive coverage of coding regions, with additional content in clinically relevant non-coding regions.
Detailed Description
Operating Principle
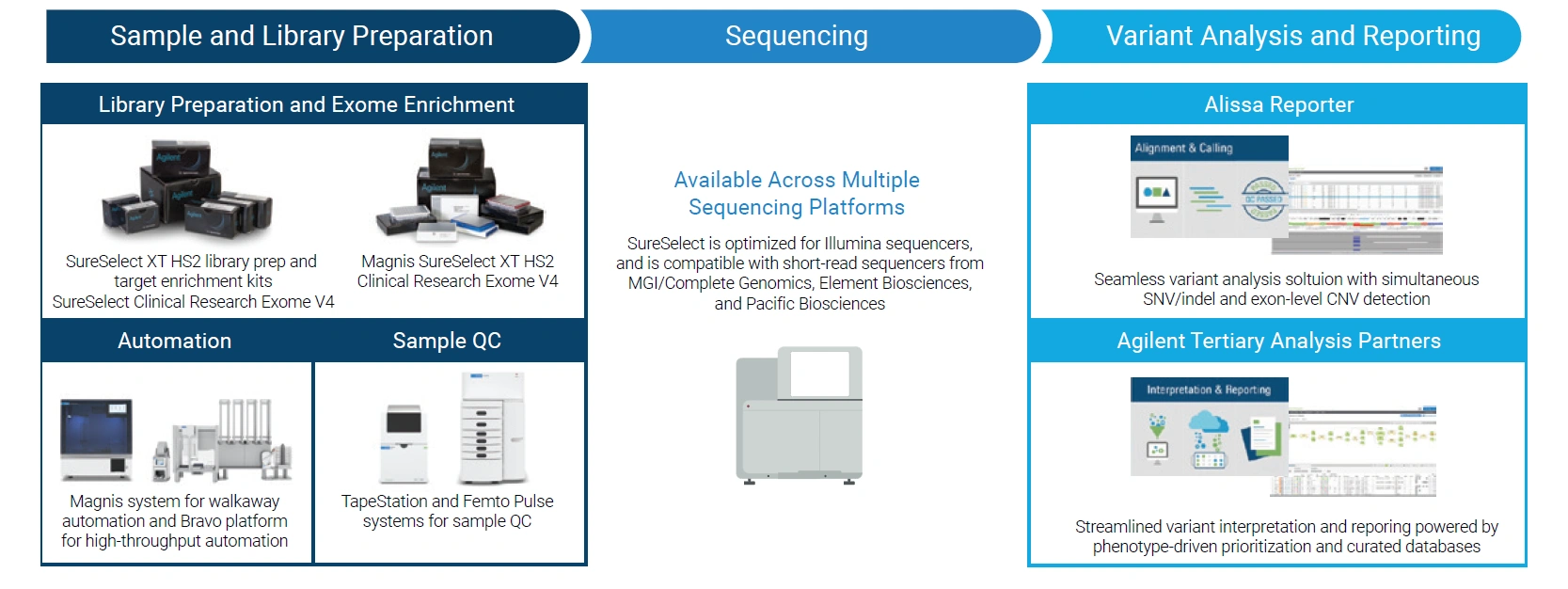
SureSelect Clinical Research Exome V4 is based on library preparation and exome capture enrichment, combining broad coverage of human coding regions with enhanced content in clinically relevant non-coding regions. Its design incorporates genomic findings and includes curated deep intronic sites, mini-genomes, and additional content associated with disease and clinical applications.
The workflow is compatible with the SureSelect XT HS2 library preparation and enrichment system.
Clinical Applications
Geared towards the identification of causal variants of genetically based diseases in selected coding and non-coding regions.
Benefits
Key Results or Indicators
Technology Used
The NGS workflow relies on SureSelect XT HS2 for library preparation and capture enrichment.
Intended Audience / User
This product is intended for clinical genetics and molecular diagnostics laboratories that require a complete workflow, from sample to result or report generation, with extensive automation capabilities.
Key Features
Ordering Information
SureSelect XT HS Clinical Research Exome V4
| 5280-0020 | 16 reactions |
| 5280-0021 | 96 reactions |
| 5280-0022 | 96 reactions Auto |
SureSelect XT PreCap Clinical Research Exome V4
| 5280-0029 | 2 hybridizations |
| 5280-0030 | 12 hybridizations |
| 5280-0031 | 12 hybridizations Auto |
Magnis SureSelect XT HS2 DNA CRE V4 ILM
| G9775A | 32 reactions |
| G9775B | 96 reactions |