Solutions for the study of genetic alterations associated with different hereditary cancer predisposition syndromes, based on next-generation sequencing and automated analysis using the SOPHiA DDM™ software.
Detailed Description
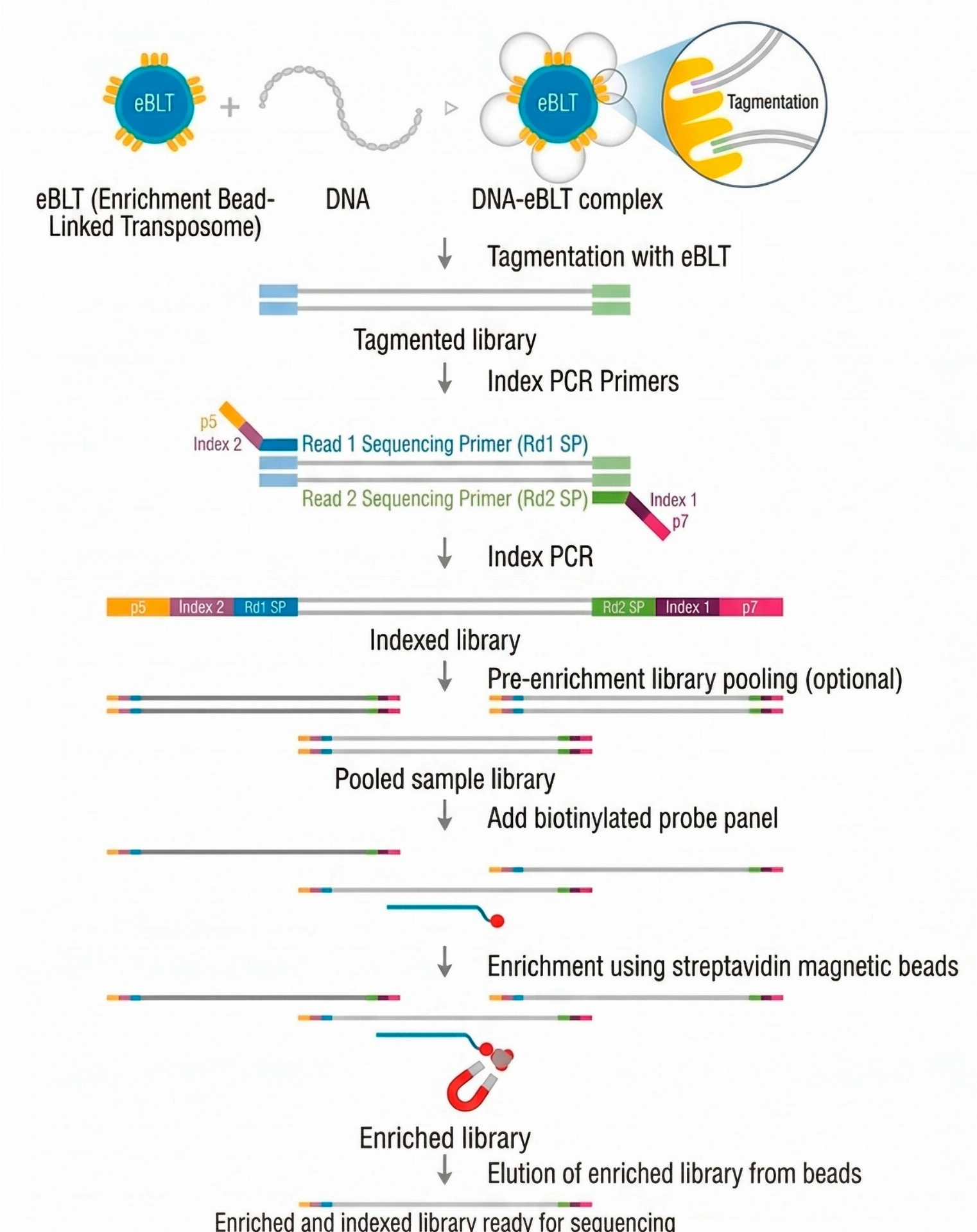
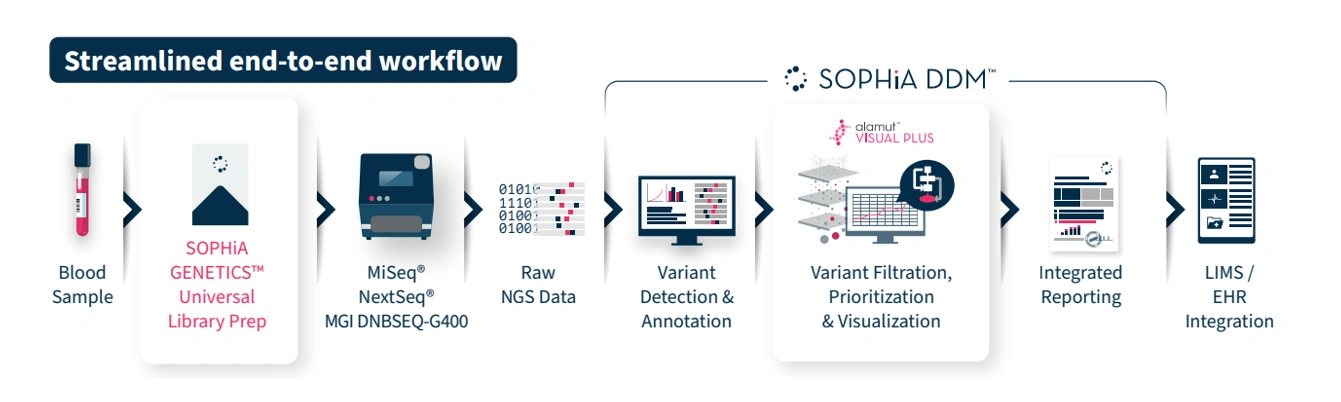
SOPHiA GENETICS™️ solutions for hereditary cancer use capture-based NGS technology to achieve uniform coverage of target regions and high on-target rates. The analytical workflow relies on SOPHiA DDM and Alamut Visual Plus, enabling the detection of complex genomic variants and efficient data interpretation.
This combination achieves high-quality variant calling and annotation, pre-classification of pathogenicity based on ACMG guidelines, and filtering/prioritization tools leading up to the generation of the final report.
Available Panels
SOPHiA GENETICS™️ Comprehensive HCS_117 Community Panel
The panel analyzes a total of 117 genes associated with hereditary cancer predisposition:
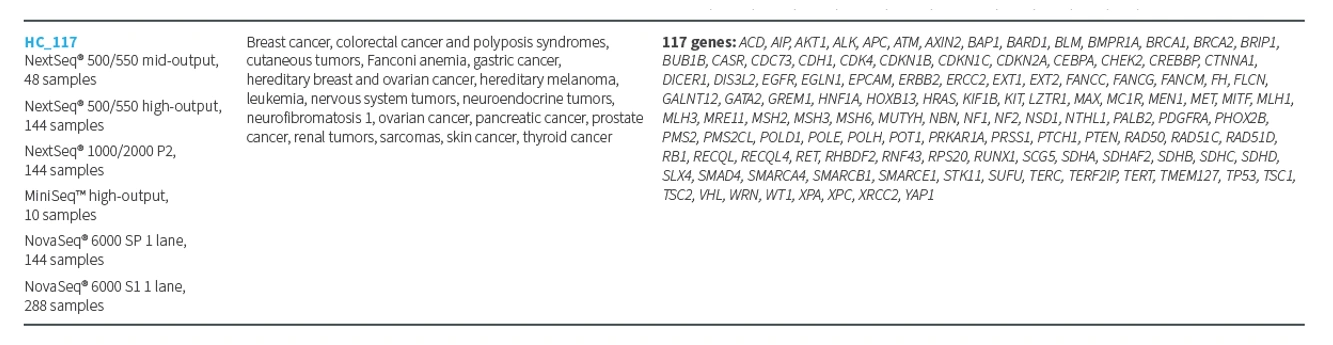

SOPHiA DDM™ Hereditary Cancer Solution (HCS) v2.0
The panel includes a total of 83 biologically relevant genes associated with major cancer predisposition syndromes, along with reinforced areas to sequence promoter regions in the APC, BRCA1, BRCA2, FAM157A, GREM1, MLH1, NTHL1, PTEN, SPINK1, and TERT genes.
Additionally, the panel features specific probes to capture SNPs that enable sample traceability.
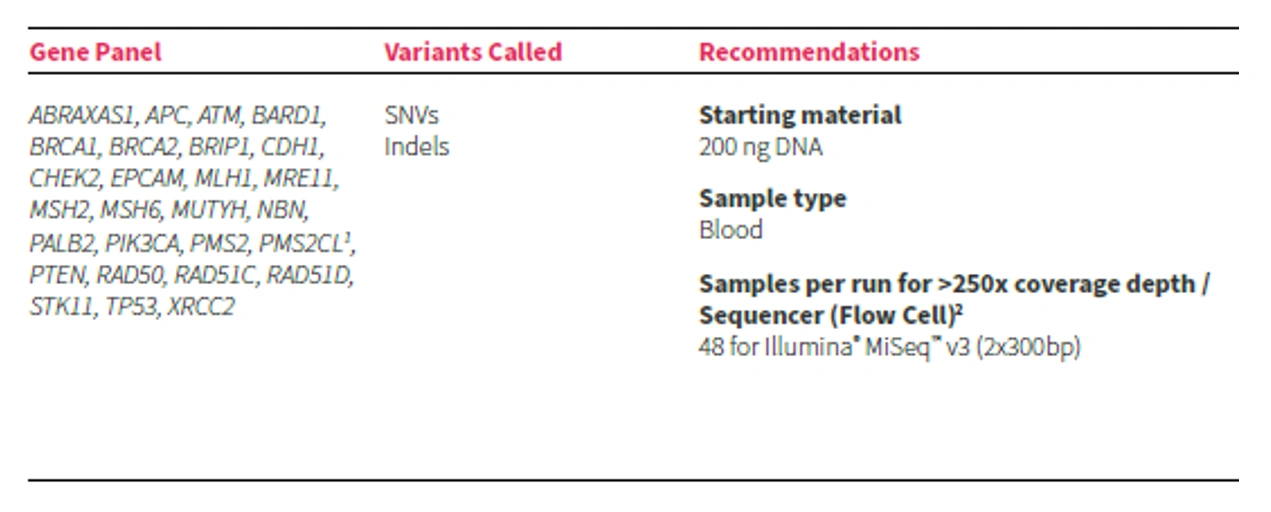
SOPHiA DDM™ Dx Hereditary Cancer Solution (HCS)
The CE-IVD marked panel includes 26 genes + 1 pseudogene:

Operating Principle
These kits are based on targeted capture enrichment techniques and subsequent sequencing on various NGS instruments. Secondary and tertiary analysis is performed on the SOPHiA DDM™ platform, which enables the detection of all types of genomic variants (SNVs, indels, and CNVs) with highly efficient clinical performance.
Clinical Applications
Hereditary cancer panel solutions are fundamental tools in clinical practice for identifying genetic alterations associated with oncology predisposition syndromes. These solutions enable efficient and precise detection of the most relevant variants, facilitating rapid diagnoses and informed therapeutic decision-making.
By focusing on key mutations related to various types of cancer, the panels offer a cost-effective and highly specific option to personalize patient prevention, monitoring, and treatment, thereby improving clinical outcomes.
Benefits
Intended Audience / User
Professional use to support healthcare professionals in informing clinical decisions related to cancer predisposition syndromes, aiding in the diagnosis and genetic counseling of familial cases.









