Kit for the determination of the GAA triplet repeats number of the FXN gene by fluorescent fragment analysis.
Information about the product
Friedreich’s ataxia (FRDA) is the most common hereditary ataxia and it´s an autosomal recessive degenerative disease.
The most common DNA abnormality associated with Friedreich’s ataxia (FRDA) is the expansion of a GAA triplet repeat polymorphism localized in the first intron of the gene encoding frataxin (FXN). Pathogenic GAA expansion alleles are in the size range of 50 to >1300 repeats with three different intervals:
- healthy (between 5-30 repeats),
- Mild symptoms (30-49 repeats)
- Severe symptoms (50-1300 repeats)
Intended Use
Adellgene® Friedreich’s Ataxia is a semi-automated in vitro diagnostic kit designed for use in clinical laboratories which quantitatively determines the number of repetitions of GAA (guanine-adenine-adenine) in the first intron of the gene encoding frataxin (FXN) located in chromosome 9 resulting in Friedreich’s ataxia disease. It aims to aid diagnosis associated with clinical findings in Friedreich’s ataxia that span from mild to severe symptoms.
The use of this kit is the determination of healthy alleles who have between 5 to 30 GAA repeats, patients with mild phenotype (30-49 repeats), and severe (50-1300).
The technology is based on the polymerase chain reaction (PCR) of genomic DNA extracted from peripheral blood followed by fluorescence analysis of the size of the PCR fragments obtained by genetic analyser and conversion of that size in the number of GAA repeats.
Patients who can benefit from this determination are those referred by a specialist. The intended user of the kit is technical personnel trained to carry out the protocol and the interpretation of results described in the instructions for use.
Workflow

Results
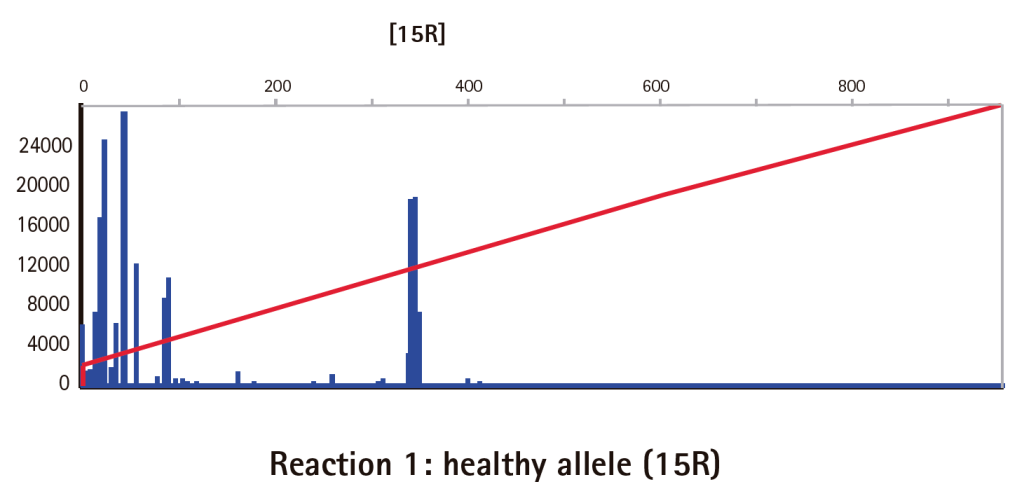
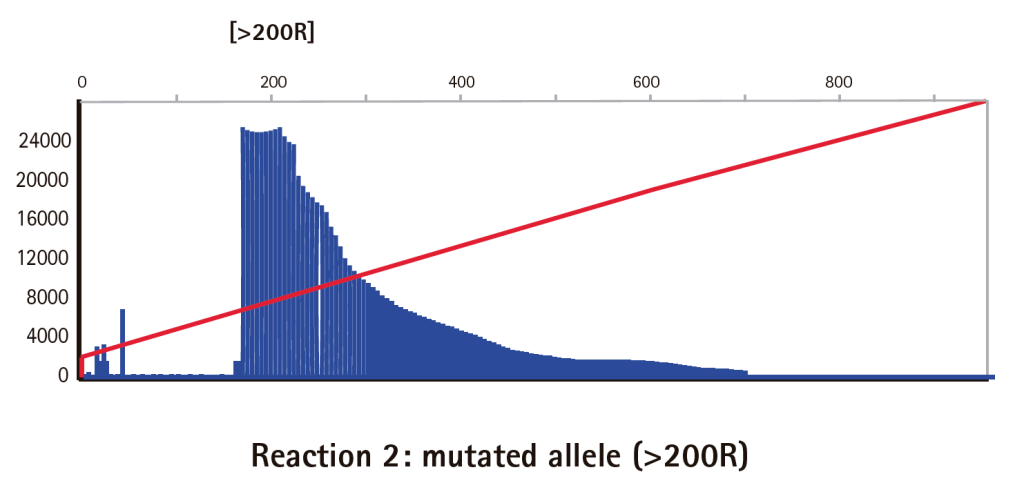

Limitations
- Mutations (point mutations, insertions, deletions) at amplification primer sites are possible and may result in the lack of allele definition. Other technologies could be necessary to resolve the genotyping.
- Data and result interpretation should be revised by qualified personnel.