IVDR workflow for whole exome sequencing (WES) that integrates Twist’s exclusive double-stranded DNA (dsDNA) probe technology with its library preparation and targeted capture reagents. The result: achieving the best coverage uniformity on the market and superior quality data for clinical environments.
Detailed Description
Operating Principle
The system combines a streamlined library preparation, based on single-tube enzymatic fragmentation, with selective enrichment of regions of interest using high-precision capture probes. The panel design ensures a high on-target rate and uniform coverage, generating highly complex libraries and homogeneous enrichment of the regions of interest.

Clinical Applications
Whole exome sequencing (WES) is currently an essential tool in clinical practice for identifying genomic alterations with diagnostic and/or therapeutic relevance. Compared to whole genome sequencing, it offers lower cost and optimal coverage of the coding regions of the human genome, facilitating more precise diagnoses and informed medical decision-making.
Benefits
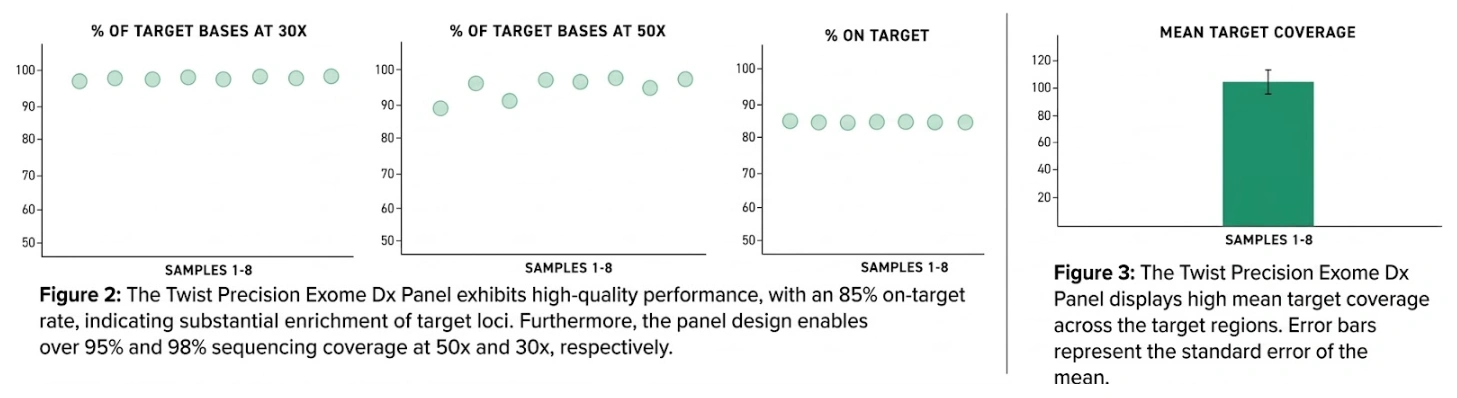
The panel captures 37.7 Mb of the human genome and includes over 97% of the reference bases such as CCDS, RefSeq, Gencode, and Ensembl v101. Additionally, it covers clinically relevant regions, such as pathogenic or likely pathogenic intronic variants reported in ClinVar, and incorporates the mitochondrial genome.
This translates into highly complex libraries, uniform enrichment, and robust performance that meets the highest IVDR standards.
Technology Used
The kits include individual library preparation reagents by enzymatic fragmentation, using Unique Dual Indices; and the necessary reagents for target enrichment by capture, including the human mitochondrial genome.
The IVDR marking guarantees reproducible results, validated quality, and tested performance.
Intended Audience/User
Molecular diagnostic laboratories working with human genomic DNA samples for whole exome applications, in in vitro diagnostic environments.
Additional Considerations
To facilitate the integration of IVDR workflows in laboratories, Twist has partnered with Platomics and its PlatoX® IVD Assistant platform, although the end-user must perform additional validation when integrating the products into diagnostic applications.
Key Features
Presentation Details
Available in two formats:
Included components (depending on kit):
References
Twist Precision Exome Dx Kits
| 104984 | 16×2 samples |
| 104985 | 96×12 samples (Plate A) |
| 104986 | 96×12 samples (Plate B) |
| 104987 | 96×12 samples (Plate C) |
| 104988 | 96×12 samples (Plate D) |
Twist Precision Prep and Enrichment Dx Kits
| 107626 | 16×2 reactions |
| 107627 | 96×12 reactions (Plate A) |
| 107628 | 96×12 reactions (Plate B) |
| 107629 | 96×12 reactions (Plate C) |
| 107630 | 96×12 reactions (Plate D) |
Twist Precision Exome Dx Panels
| 107619 | 2 reactions |
| 107620 | 12 reactions |









