PGTA-ChromInst® (Preimplantation Genetic Testing for Aneuploidies) is a PGT-A solution. It combines in a single step WGA (Whole Genome Amplification) and library preparation for NGS (Next Generation Sequencing), (MALBAC® technology), guaranteeing a uniform and efficient amplification of DNA (MALBAC® technology).

MALBAC® technology
WGA methods are prone to introduce amplification bias, as they amplify some regions better than others. However, MALBAC® (Multiple Annealing and Looping Based Amplification Cycles) technology results in amplification products that have complementary ends capable of forming loops of DNA, which enhances the quasi-linearity of amplification. In this way, much better amplification uniformity is achieved than with the other WGA methods described to date.
High genome coverage and amplification of >90% of loci is achieved with this technology. In addition, the ADO (Allele Drop Out) rate is <10%.
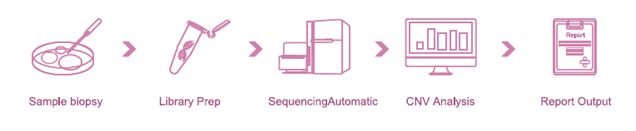
Features:
- Easy:
<1.5hrs hand-on work; WGA and Lib Prep are combined in one step.
- Safe:
Single tube/well reactions to avoid sample loss and contamination.
- Timesaving:
The whole process can be finished within 24 hours (9 hours if coupled with Illumina’s MiniSeq or MiSeq sequencers, and the auto-analysis bioinformatics platform ChromGO™).
- Flexible:
Compatible with both Illumina (Miniseq and MiSeq included) and Ion Torrent sequencing platforms.
- Accurate:
Most advanced single cell WGA technique.
- Complete:
An entire solution, from sample preparation to report output.
ChromGO™
Simple data analysis solution, which allows you to quickly customize your data analysis reports on both online and local servers.
Features:
- Intuitive and simple interface.
- Versatile: data can be analyzed with different criteria (mosaicism, gender information…).
- Detailed: details QC (Quality Control), including reads, GC content and CV (coefficient variation) presented in individual files.
- Flexible: NGS data from different sequencing platforms can be processed.
- 1M reads/sample for the detection of 10Mb deletions.

Who is it for?
- Women with advanced maternal age.
- Men with azoospermia.
- Patients with recurrent pregnancy losses.
- Patients with repeated implantation failures.
- Patients with chromosome structural abnormalities, such as reciprocal translocation and Robertsonian translocation.