Kit for detecting the S65C mutation of HFE gene by Real Time PCR using TaqMan® probes technology
Information about the product
Hereditary hemochromatosis (HH) is an autosomal recessive inherited disorder of iron metabolism. Due to excessive intestinal absorption, iron accumulates in the parenchymal cells of the liver, pancreas, heart and other organs resulting in damage to its structure and its function impaired. Although the disease symptoms are often non-specific, much of the organ damage is irreversible once it has occurred. Early detection and treatment is therefore very important as part of preventive medicine.
A number of different HFE mutations have been described. One of these mutations is a substitution at position 193 (A -> T) of exon 2 resulting in an amino acid change at position 65 from serine to cysteine (S65C) and has proven to be generally benign, although the C282Y/S65C genotype can impart a slight increase in disease risk, contributing to a mild disease phenotype.
Intended Use
Genvinset® HFE S65C is a semi-automated in vitro diagnostic kit for the qualitative detection of the S65C mutation (NCBI dbSNP rs1800730; NM_000410.4:c.193A>T), in the HFE gene (OMIM: 613609) associated with primary hemochromatosis, in genomic DNA extracted from whole blood using Real Time PCR technology with specific TaqMan® probes.
Patients who can benefit from this determination are those referred by a specialist. The results of this test should not be the only ones on which the therapeutic decision is based and should be used as an aid in the diagnosis together with results of other markers of the disease.
The intended user of the kit is technical personnel trained to carry out the protocol and the interpretation of results described in the instructions for use.
Workflow

Results
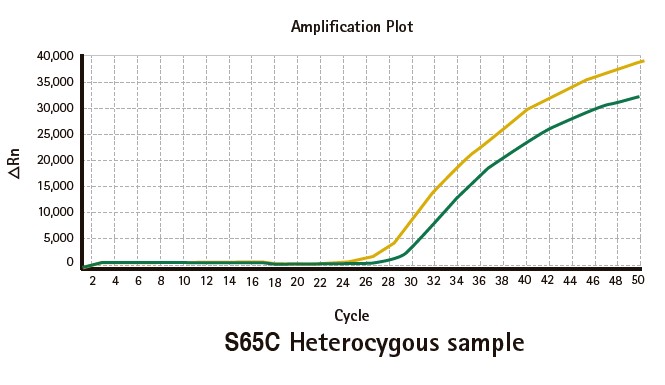
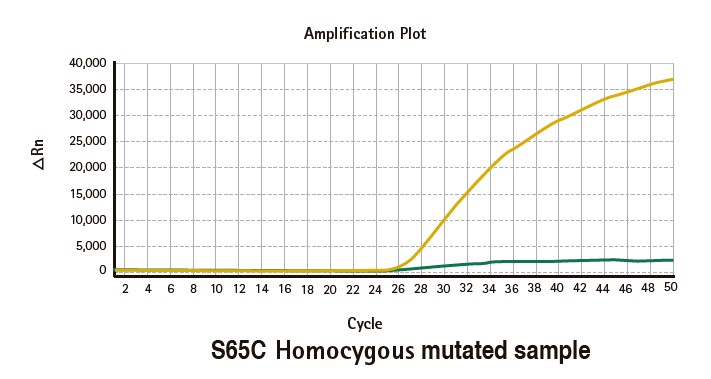
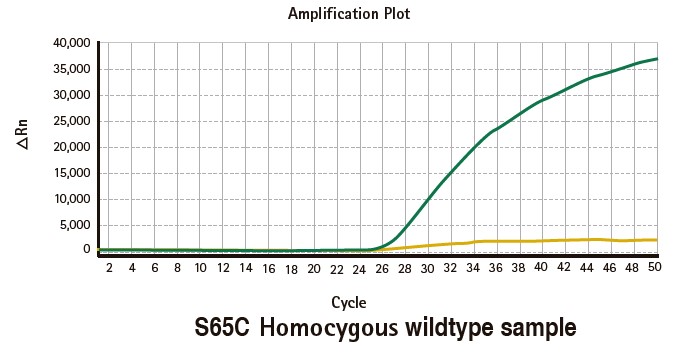

Limitations
- Mutations or polymorphisms at annealing primer/probe sites are possible and may result in the lack of allele definition. Other technologies could be necessary to resolve the typing.
- Data and result interpretation should be revised by qualified personnel.
- This product is an auxiliary tool for the diagnosis of patients with suspected hereditary hemochromatosis. Use these results in conjunction with clinical data and results of other tests performed on the patient.