Spike-in panels designed to complement exome sequencing through evenly distributed genome-wide coverage, aimed at improving the detection of copy number variations (CNVs). Available in three probe densities—100 kb, 50 kb, and 25 kb—they seamlessly integrate into standard Twist target enrichment and capture workflows.
Detailed Description
Operating Principle
Exome sequencing focuses on coding regions and other specific areas of interest, which typically represent only about 1–2% of the genome. This limitation leaves large intergenic regions out of the analysis and hampers CNV detection when the exome is used on its own.
Twist CNV Backbone Spike-in panels add a series of probes distributed at regular intervals across the genome to improve resolution in CNV calling. These probes target common and polymorphic SNPs across multiple populations and are spaced at 25 kb, 50 kb, or 100 kb, depending on the resolution required by each laboratory.
Clinical Applications
Designed for cytogenetic research, this panel expands the scope of exome sequencing in the analysis of samples with clinically relevant chromosomal alterations. It facilitates genome-wide CNV detection, including large events that traditionally required techniques such as CMA or aCGH. Furthermore, it adds value to comprehensive variant studies by enabling the simultaneous analysis of CNVs, LOH, SNVs, and small indels within a single NGS approach.
Benefits
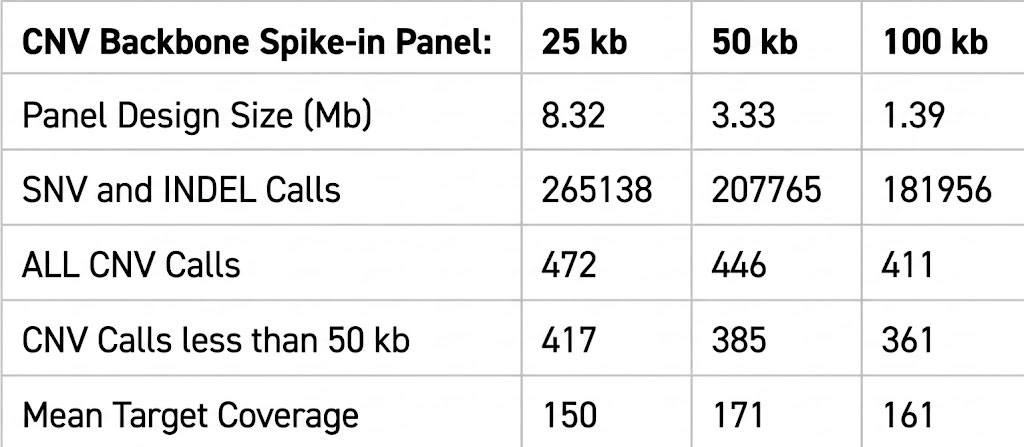
Intended Audience / User
Key Features
Ordering Information
Commercial Configurations
| 2-reaction kit | (16 samples) |
| 12-reaction kit | (96 samples) |
Available Resolutions
| 110756 / 110757 | 25 kb: high resolution (8.32 Mb) |
| 110758 / 110759 | 50 kb: intermediate resolution (3.33 Mb) |
| 110760 / 110761 | 100 kb: lower resolution (1.39 Mb) |