Kit para la determinación de alelos sanos y premutados del gen FMR1 mediante análisis de fragmentos fluorescentes.
Información del producto
El síndrome del X Frágil (FXS, OMIM # 300624) es una enfermedad ligada al cromosoma X, basada fundamentalmente en la expansión genómica de un triplete de nucleótidos (CGG), y en una metilación aberrante de la región promotora.
EL FXS tiene una prevalencia de 1 de cada 4.000 hombres y 1 de cada 8.000 mujeres. Los individuos afectados muestran n fenotipo llamativo que consiste en orejas grandes y una mandíbula prominente.
En función del número de repeticiones de este triplete, se pueden establecer tres categorías:
- Hasta 45 o 55 repeticiones: individuos con alelos sanos.
- De 45 o 55 a 200 repeticiones: individuos con alelos premutados.
- Más de 200 repeticiones: individuos con alelos mutantes.
Uso previsto
Adellgene® Fragile X Screening es un kit de diagnóstico in vitro semiautomatizado de uso en laboratorios clínicos diseñado para la determinación cuantitativa del número de repeticiones del triplete CGG (citosina-guanina-guanina) presentes en la region no traducida 5’ del gen que ocasiona retraso mental por X frágil (FMR1, del inglés “Fragile X mental retardation-1”). Esta determinación pretende ayudar en el diagnóstico clínico asociado con el síndrome X Frágil, por ejemplo, retraso mental, fallo ovárico primario, y temblores/ataxia.
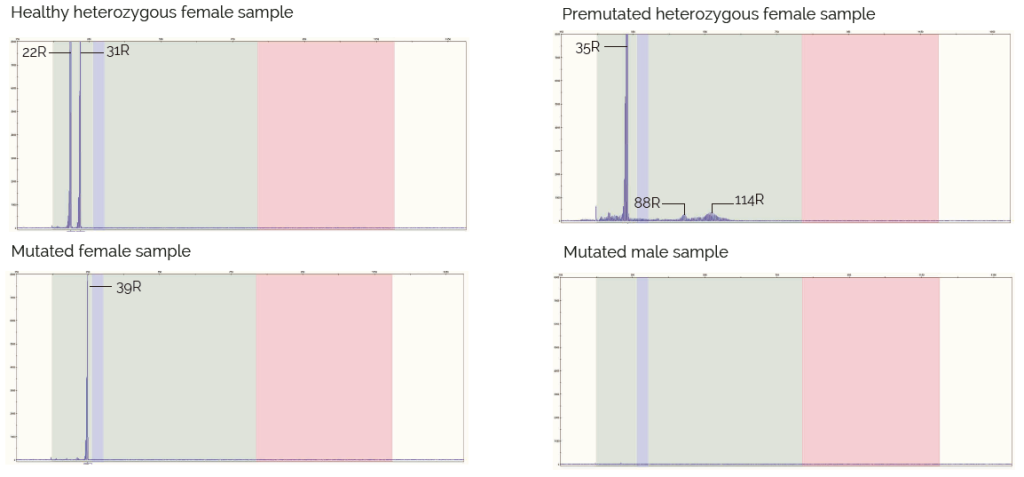
Este kit se puede utilizar para la determinación del número de repeticiones presentes en individuos sanos (hasta 45/55 repeticiones CGG) e individuos premmutados (de 45/55 a 200 repeticiones de tripletes CGG). Las muestras de mujer que reportan solo un pico en el electroferograma y las muestras de varón sin pico deben analizarse utilizando otra técnica apropiada, por ejemplo, TP-PCR, para verificar la inexistencia de un alelo completamente mutado (repeticiones de >200 CGG).
La determinación se basa en la amplificación del ADN genómico extraído a partir de sangre periférica mediante reacción en cadena de polimerasa (PCR), posterior análisis del tamaño de los fragmentos fluorescentes amplificados en un analizador genético y conversión del tamaño del fragmento en su respectivo número de repeticiones CGG.
Los pacientes que pueden beneficiarse de esta determinación son los remitidos por un especialista. El usuario previsto de este kit es personal técnico entrenado y cualificado para realizar el protocolo descrito en las instrucciones de uso y en la interpretación de sus resultados.
Flujo de trabajo

Resultados
Limitaciones
- Las mutaciones (mutaciones puntuales, inserciones, deleciones) en los sitios de hibridación de los primers pueden dar lugar a la falta de definición de alelos. Otras tecnologías podrían ser necesarias para resolver el genotipado.
- Los resultados homocigotos y los de varones en los que no se detecta ningún alelo, deben ser confirmados por procedimientos alternativos.
- Los datos y la interpretación de los resultados deben ser revisados por personal cualificado.