Information about the product
Susceptibility to gluten sensitivity is, in part, genetically determined. The strong predisposition is associated with HLA-DQ alleles, encoding the α and ß chains of two molecules of the Major Histocompatibility Complex (MHC) class II.
In most populations studied, 90-95% of patients carry the HLA-DQ2 heterodimer encoded by the alleles DQA1*05 and DQB1*02 in cis position or in trans position. The remaining patients (5-10%) usually carry a second heterodimer, HLADQ8 (majority among South American indigenous patients), encoded by the alleles DQA1*03:01 and DQB1*03:02. It is estimated that only 0.5% of celiac patients aren’t DQ2 or DQ8.
Intended Use
Genvinset® HLA Celiac is a semi-automated in vitro diagnostic kit for the qualitative detection of the HLA-DQB1*02, DQB1*03:02 and DQA1*05 alleles in genomic DNA extracted from whole blood, and the consequent determination of the DQ2 and DQ8 antigens, associated with celiac disease predisposition. The kit is able to determine the homozygosity or heterozygosity status for the DQB1*02 alleles. The analysis is based on real-time PCR technology, using TaqMan® probes.
Patients who can benefit from this determination are those referred by a specialist. The results of this test should not be the only ones on which the therapeutic decision is based and should be used as an aid in the diagnosis together with results of other markers of the disease.
The intended user of the kit is technical personnel trained to carry out the protocol and the interpretation of results described in the instructions for use.
Workflow

Results
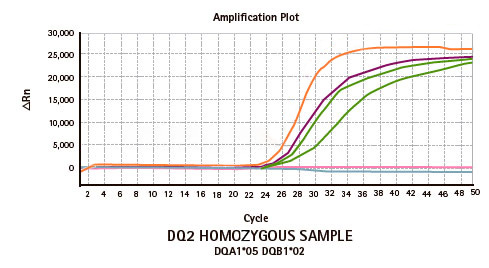
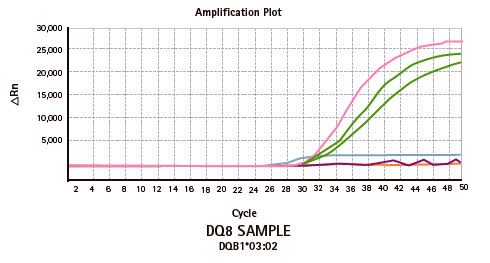
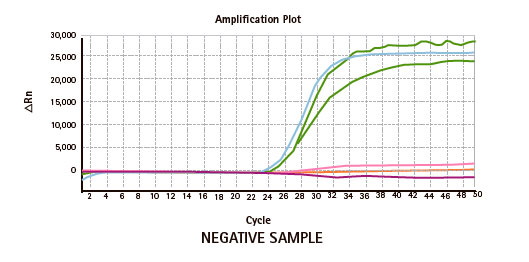
Limitations
- Mutations or polymorphisms at annealing primer/probe sites are possible and may result in the lack of allele definition. Other technologies could be necessary to resolve the typing.
- Data and result interpretation should be revised by qualified personnel.
- This product is an auxiliary tool for the diagnosis of patients with suspected celiac disease. Use these results in conjunction with clinical data and results of other tests performed on the patient.