Digital PCR assay for the precise quantification of donor-derived cell-free DNA (dd-cfDNA) in transplant patients.
Detailed Description
HoloGRAFT ONE is a digital PCR (dPCR) assay developed by Werfen for the detection and quantification of donor-derived cell-free DNA (dd-cfDNA) in the plasma of transplant recipient patients. This molecular marker has established itself as a useful tool for non-invasive monitoring of graft status.
During the process of cell damage in the graft—whether due to rejection, infection, or ischemic damage—donor DNA fragments are released into the recipient’s bloodstream. HoloGRAFT ONE allows for the specific detection and quantification of these fragments using digital PCR, a technology that offers superior sensitivity and precision compared to other molecular methods.
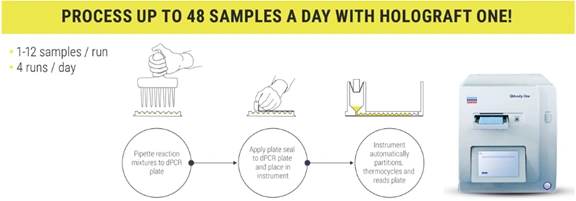
The methodology requires DNA extraction from plasma, followed by a dPCR reaction with specific probes previously selected based on the genetic differences between donor and recipient. The assay measures the percentage of dd-cfDNA relative to total cell-free DNA, expressed as a percentage.
This value allows the clinician to identify graft injury events early, facilitating therapeutic decisions based on an objective biomarker. HoloGRAFT ONE is applicable to various types of solid organ transplants, such as kidney, liver, heart, or lung, and is easily integrated into post-transplant clinical follow-up workflows.
Key Features