High-resolution NGS-based HLA typing system that supports multiple sequencing platforms and offers maximum workflow flexibility.
Detailed Description
HLA-SuBiTo NGStype® is a comprehensive solution for high-resolution HLA typing based on next-generation sequencing (NGS), developed by inno-train Diagnostik. The system allows the simultaneous analysis of up to 11 HLA loci using a long-range PCR approach, compatible with multiple library preparation methods and sequencers.
The technology adapts to both short- and long-read sequencers, including Illumina, Oxford Nanopore Technologies (ONT), and MGI. This gives the user complete freedom to choose the sequencing platform according to their infrastructure or analysis needs.
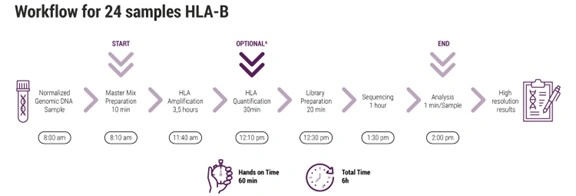
The workflow begins with a long-range PCR covering the HLA-A, -B, -C, -DPA1, -DPB1, -DQA1, -DQB1, and DRB1/3/4/5 loci, with amplicons ranging from 3.4 to 7.4 kb. Analysis quality is guaranteed by the use of specific primers and an optimized master mix included in the kit.
Depending on the chosen workflow, the process can be adapted for laboratories analyzing from a single sample (using ONT’s MinION) up to hundreds of samples in parallel (using MiSeq, MiniSeq, iSeq, among others). Flexibility is one of its main advantages, along with the system’s scalability.
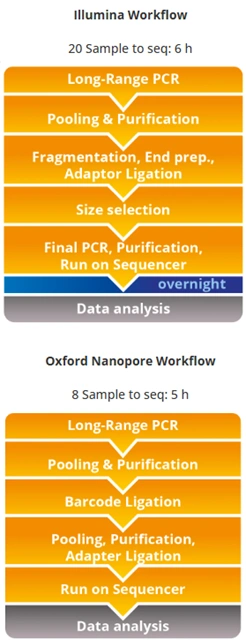
Library preparation can be performed with validated third-party kits such as NEBNext UltraExpress™ for Illumina or Rapid Barcoding Kit for ONT, and the launch of a proprietary library preparation kit by inno-train is anticipated.
Data analysis is performed using NGStype® software, compatible with fastq files from any platform. This software includes optimized algorithms for short- and long-read sequences, allowing automatic haplotype analysis and clear visualization of results.