Comprehensive solution for chimerism study by qPCR, combining pre-transplant genetic typing kits and post-transplant quantitative monitoring.
Detailed Description
KMRtype® and KMRtrack® are complementary systems developed by GenDx for the characterization and monitoring of chimerism in transplant patients, using qPCR-based assays. The system allows everything from the identification of unique informative markers between recipient and donor, to the precise quantification of reference DNA in post-transplant samples.
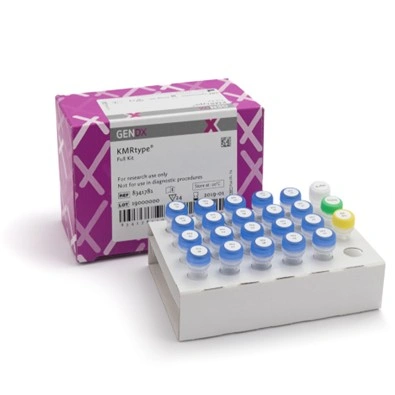
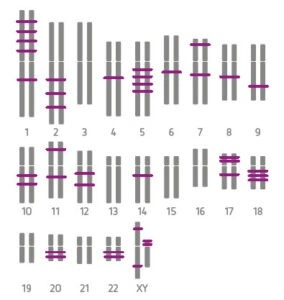
KMRtype® is a genetic typing kit that allows the selection of informative markers between recipient and donor DNA. Each mix contains three multiplexed primers and is used in pre-transplant settings. The kits are available in Core (30 assays in 10 mixes) and Extended (15 assays in 5 additional mixes) versions, allowing coverage of up to 45 assays in total. This phase is essential to identify which specific markers can be used effectively for monitoring.
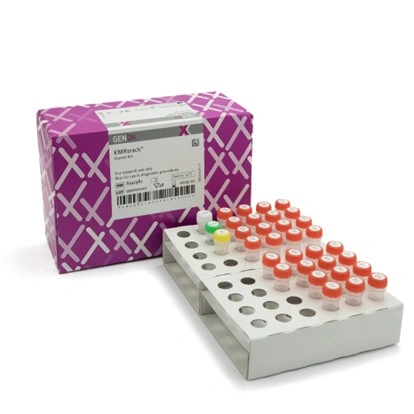
KMRtrack® is used for post-transplant monitoring using singleplex qPCR assays. Each marker selected in the typing phase is used to quantify the proportion of reference DNA (either from the recipient or the donor) in the samples, allowing precise monitoring of the patient’s chimeric status over time. The achieved limit of detection is 0.05%, depending on the amount of DNA used.
Both kits are optimized for use alongside the KMRengine® software, which guides experimental setup, manages qPCR plates, analyzes results, and automatically reports the chimerism percentage. The platform allows registering multiple samples, designing customized experiments, and visualizing data graphically and chronologically.
Performance and Validation
Validation studies showed 100% accuracy in typing and high intra- and inter-batch reproducibility in monitoring. Specificity was confirmed for all markers. Detection tests indicated that the system maintains robust performance with variable amounts of DNA, making it ideal for both clinical and experimental samples.
Important Considerations
Clinical Applications
This system is indicated for chimerism studies in the context of hematopoietic stem cell transplantation, with special utility in graft monitoring and early detection of relapse. Its sensitivity allows the detection of minimal amounts of donor or recipient-derived DNA, facilitating early therapeutic decisions.